Services
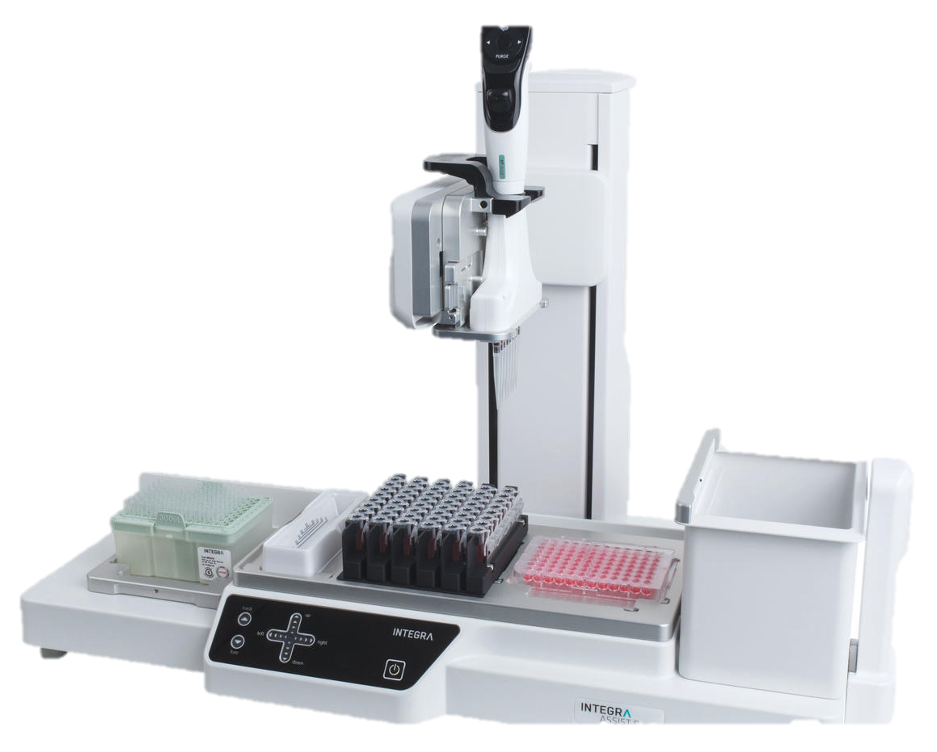
Services provided by the genomics core facility include DNA and RNA quantification, chemical and mechanical fragmentation, NextGen sequencing library preparation and normalization, whole genome de novo and resequencing, whole transcriptome sequencing (RNASeq), and whole exome sequencing. We also design specialized library prep and seq methods such as whole genome methylation analysis and targeted region of interest sequencing.
Instrumentation
The NextSeq 500 is the sequencer of choice for any size of projects. This tabletop
sequencing system is capable of sequencing a high coverage (30x) whole human genome
in one run. Its flexible throughput means no waiting to fill a flowcell and thus a
quicker turnaround time for your sequencing sample. Because of high- and mid-output
options, the NextSeq 500 can easily shift between high- and low-throughput sequencing
projects. The NextSeq 500 generates 130-800 million sequence reads in 75-300 bp read
length depending on the selected run mode. It is operated by the Genomics Core staff
to process the user-submitted samples as well as those prepared by the facility.
.
The MiSeq desktop sequencer has broad applications in targeted gene sequencing, metagenomics, small genome and transcriptome sequencing, targeted gene expression, and amplicon sequencing. The MiSeq system generates 1-25 million reads in 35-600 bp read length in multiple run modes, which makes the system versatile enough to provide sequencing capacity for both small pilot studies to full-scale genomic experiments. The system is operated by the Genomics Core staff to process the user-submitted samples as well as those prepared by the facility.

The NanoDrop One is available for measuring concentration of nucleic acids and protein samples. and assessing purity. The full spectra (190 nm to 850 nm) spanning the ultraviolet - visible range is obtained within seconds using 1-2 microliters of samples. The NanoDrop is free of charge and available for the users trained by the Genomics Center staff.
The Qubit has the ability to differentiate between dsDNA, ssDNA, RNA, and protein. Concentration is proportional to the fluorescence signal from the dye that binds specifically to the nucleic acid or protein sample being assayed. Most contaminants have no effect on the final result. This makes the Qubit extremely useful when accurate quantitation is needed for downstream applications, such as next-generation sequencing experiments. The Qubit is available for the trained users at the cost of the consumable.

The Agilent 4200 TapeStation system is an automated gel electrophoresis system that can process 1-96 samples per run with minimum hands-on time. Many sample types can be analyzed on the TapeStation, including total RNA, labeled RNA, small and micro RNAs, as well as small and large DNA fragments. The TapeStation allows for a constant cost regardless of the number of samples. Users are no longer forced to pay in 12 sample increments. The TapeStation is available for the trained users at the cost of the consumable.